Difference between revisions of "DivIVA"
(→Expression and regulation) |
(→References) |
||
| Line 125: | Line 125: | ||
=References= | =References= | ||
| − | + | ==Reviews== | |
| − | <pubmed>19654604,19666580,9219999,19019154,15554965, 12368265,11445541,10835369,12511520,14651647, 19478798 ,19429628, 11445541, 9219999, 9045828 </pubmed> | + | <pubmed> 19654604, 19884039 </pubmed> |
| − | + | ==Original Publications== | |
| + | <pubmed>19654604, 19666580,9219999,19019154,15554965, 12368265,11445541,10835369,12511520,14651647, 19478798 ,19429628, 11445541, 9219999, 9045828 </pubmed> | ||
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 20:24, 4 November 2009
- Description: cell-division initiation protein (septum placement)
| Gene name | divIVA |
| Synonyms | ylmJ |
| Essential | no |
| Product | cell-division initiation protein |
| Function | septum placement |
| MW, pI | 19 kDa, 4.846 |
| Gene length, protein length | 492 bp, 164 aa |
| Immediate neighbours | ylmH, ileS |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 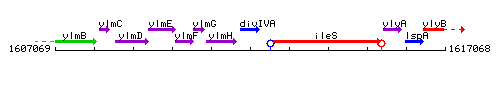
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU15420
Phenotypes of a mutant
Deletion of divIVA leads to filamentation and polar divisions that in turn cause a minicell phenotype. PubMed A divIVA mutant has a severe sporulation defect. PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
Filamentation is suppressed by mutations in minCD PubMed.
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: DivIVA is required for polar localisation of MinCD via MinJ. PubMed It also recruits RacA to the distal pole of the prespore PubMed.
- Protein family: gpsB family (according to Swiss-Prot)
- Paralogous protein(s): GpsB
Extended information on the protein
- Kinetic information:
- Domains: The first 60 amino acids constitute a lipid binding domain. PubMed
Multimerisation involves two coiled-coil motifs, one in the lipid binding domain, and the other one being present in the helical C-terminal domain PubMed.
- Modification: The Mycobacterium DivIVA homologue Wag31 is phosphorylated at T73 PubMed.
- Cofactor(s): not known
- Effectors of protein activity: not known
- Localization: DivIVA forms a ring underneath the invaginating membrane at the site of cell division and is enriched at both cell poles PubMed.
Database entries
- Structure:
- UniProt: P71021
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Operon: divIVA PubMed
- Additional information:
Biological materials
- Mutant: divIVA::tet available from the Hamoen Lab
- Expression vector:
- lacZ fusion:
- GFP fusion: divIVA-gfp fusions available from the Hamoen Lab
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Leendert Hamoen, Centre for Bacterial Cell Biology, Newcastle upon Tyne, United Kingdom [4]
Imrich Barak, Slovak Academy of Science, Bratislava, Slovakia homepage
Your additional remarks
References
Reviews
Marc Bramkamp, Suey van Baarle
Division site selection in rod-shaped bacteria.
Curr Opin Microbiol: 2009, 12(6);683-8
[PubMed:19884039]
[WorldCat.org]
[DOI]
(I p)
Jennifer R Juarez, William Margolin
Irresistible curves.
EMBO J: 2009, 28(15);2147-8
[PubMed:19654604]
[WorldCat.org]
[DOI]
(I p)
Original Publications