Difference between revisions of "CypC"
| Line 1: | Line 1: | ||
| − | * '''Description:''' | + | * '''Description:''' fatty acid beta-hydroxylating cytochrome P450, hydroxylates myristic acid to beta-hydroxymyristic <br/><br/> |
{| align="right" border="1" cellpadding="2" | {| align="right" border="1" cellpadding="2" | ||
Revision as of 17:39, 1 February 2010
- Description: fatty acid beta-hydroxylating cytochrome P450, hydroxylates myristic acid to beta-hydroxymyristic
| Gene name | cypC |
| Synonyms | ybdT |
| Essential | no |
| Product | fatty acid beta-hydroxylating cytochrome P450 |
| Function | biosynthesis of beta-hydroxy fatty acid |
| MW, pI | 47 kDa, 6.468 |
| Gene length, protein length | 1251 bp, 417 aa |
| Immediate neighbours | ybxI, ybyB |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 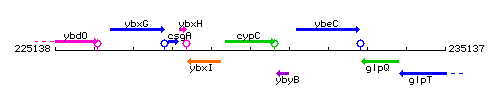
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU02100
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: cytochrome P450 family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization:
Database entries
- Structure:
- UniProt: O31440
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Operon: cypC (according to DBTBS)
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Dirk Höper, Uwe Völker, Michael Hecker
Comprehensive characterization of the contribution of individual SigB-dependent general stress genes to stress resistance of Bacillus subtilis.
J Bacteriol: 2005, 187(8);2810-26
[PubMed:15805528]
[WorldCat.org]
[DOI]
(P p)
Dong-Sun Lee, Akari Yamada, Hiroshi Sugimoto, Isamu Matsunaga, Hisashi Ogura, Kosuke Ichihara, Shin-Ichi Adachi, Sam-Yong Park, Yoshitsugu Shiro
Substrate recognition and molecular mechanism of fatty acid hydroxylation by cytochrome P450 from Bacillus subtilis. Crystallographic, spectroscopic, and mutational studies.
J Biol Chem: 2003, 278(11);9761-7
[PubMed:12519760]
[WorldCat.org]
[DOI]
(P p)
Dong Sun Lee, Akari Yamada, Isamu Matsunaga, Kosuke Ichihara, Shin-ichi Adachi, Sam-Yong Park, Yoshitsugu Shiro
Crystallization and preliminary X-ray diffraction analysis of fatty-acid hydroxylase cytochrome P450BSbeta from Bacillus subtilis.
Acta Crystallogr D Biol Crystallogr: 2002, 58(Pt 4);687-9
[PubMed:11914497]
[WorldCat.org]
[DOI]
(P p)
I Matsunaga, A Ueda, N Fujiwara, T Sumimoto, K Ichihara
Characterization of the ybdT gene product of Bacillus subtilis: novel fatty acid beta-hydroxylating cytochrome P450.
Lipids: 1999, 34(8);841-6
[PubMed:10529095]
[WorldCat.org]
[DOI]
(P p)