Difference between revisions of "GltC"
(→Basic information/ Evolution) |
(→Labs working on this gene/protein) |
||
| Line 139: | Line 139: | ||
[[Stülke|Jörg Stülke]], University of Göttingen, Germany | [[Stülke|Jörg Stülke]], University of Göttingen, Germany | ||
[http://wwwuser.gwdg.de/~genmibio/stuelke.html Homepage] | [http://wwwuser.gwdg.de/~genmibio/stuelke.html Homepage] | ||
| + | |||
| + | [[Fabian_Commichau|Fabian Commichau]] University of Göttingen, Germany | ||
| + | [http://genmibio.uni-goettingen.de/index.php?id=130 Homepage] | ||
=Your additional remarks= | =Your additional remarks= | ||
Revision as of 09:19, 28 January 2012
- Description: Transcriptional activator of the gltA-gltB operon. Activates expression of the operon in the absence of arginine.
| Gene name | gltC |
| Synonyms | |
| Essential | No |
| Product | transcriptional regulator (LysR family) |
| Function | positive regulation of the glutamate synthase operon (gltAB) |
| Interactions involving this protein in SubtInteract: GltC | |
| Metabolic function and regulation of this protein in SubtiPathways: Ammonium/ glutamate | |
| MW, pI | 33.9 kDa, 5.62 |
| Gene length, protein length | 900 bp, 300 amino acids |
| Immediate neighbours | gltA, proJ |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 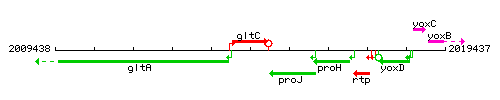
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
biosynthesis/ acquisition of amino acids, transcription factors and their control
This gene is a member of the following regulons
The GltC regulon:
The gene
Basic information
- Locus tag: BSU18460
Phenotypes of a mutant
gltC mutants are auxotrophic for glutamate.
Database entries
- DBTBS entry: [1]
- SubtiList entry:[2]
Additional information
The protein
Basic information/ Evolution
- Protein family: LysR family PubMed
- Paralogous protein(s): none, but there are 19 members of the LysR family in B. subtilis
Extended information on the protein
- Kinetic information:
- Domains: DNA-binding helix-turn-helix motif: AA 18 ... 37
- Modification:
- Cofactor(s):
- Effectors of protein activity: 2-oxoglutarate stimulates transcription activation, glutamate inhibits transcription activation PubMed
Database entries
- Structure:
- UniProt: P20668
- KEGG entry: [3]
Additional information
Expression and regulation
- Regulation: autoregulation by GltC PubMed
- Regulatory mechanism: autorepression PubMed
- Database entries: DBTBS
- Additional information:
Biological materials
- Mutant: GP344 (erm), GP738 (spc) (available in Stülke lab)
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Stülke lab
- Antibody: available in Stülke lab
Labs working on this gene/protein
Linc Sonenshein, Tufts University, Boston, MA, USA Homepage
Jörg Stülke, University of Göttingen, Germany Homepage
Fabian Commichau University of Göttingen, Germany Homepage
Your additional remarks
References
Reviews
Sabine Brantl, Andreas Licht
Characterisation of Bacillus subtilis transcriptional regulators involved in metabolic processes.
Curr Protein Pept Sci: 2010, 11(4);274-91
[PubMed:20408793]
[WorldCat.org]
[DOI]
(I p)
Original Publications