Difference between revisions of "RnjB"
| Line 37: | Line 37: | ||
=== Basic information === | === Basic information === | ||
| − | * ''' | + | * '''Locus tag:''' |
===Phenotypes of a mutant === | ===Phenotypes of a mutant === | ||
Revision as of 15:35, 2 June 2009
- Description: RNase J2
| Gene name | rnjB |
| Synonyms | ymfA |
| Essential | no |
| Product | RNase J2 |
| Function | RNA processing and degradation |
| MW, pI | 56 kDa, 9.18 |
| Gene length, protein length | 1545 bp, 515 aa |
| Immediate neighbours | dapA, tepA |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 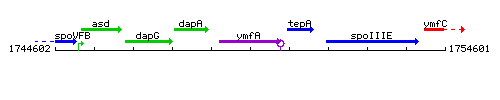
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag:
Phenotypes of a mutant
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: endoribonuclease, involved in processing of thrS mRNA
- Protein family: RNase J subfamily (according to Swiss-Prot)
- Paralogous protein(s): RnjA
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization: cytoplasm (according to Swiss-Prot)
Database entries
- Structure: 3BK1 (RNase J from Thermus thermophilus) 3BK2 (RNase J from Thermus thermophilus, complex with UMP)
- Swiss prot entry: O31760
- KEGG entry: BSU16780
- E.C. number:
Additional information
Expression and regulation
- Operon:
- Sigma factor:
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant: GP45, GP1113 (spc), available in Stülke lab
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Stülke lab
- Antibody:
Labs working on this gene/protein
Harald Putzer, IBPC Paris, France Homepage
Your additional remarks
References
- Even, S., Pellegrini, O., Zig, L., Labas, V., Vinh, J., Brechemmier-Baey, D., and Putzer, H. (2005) Ribonucleases J1 and J2: Two novel endoribonucleases in B. subtilis with functional homology to E. coli RNase E. Nucl Acids Res 33, 2141-2152. PubMed
- de la Sierra-Gallay IL, Zig L, Jamalli A, Putzer H. (2008 Structural insights into the dual activity of RNase J. Nat. Struct. Mol. Biol. 15:206-212. PubMed
- Mäder, U., Zig, L., Kretschmer, J., Homuth, G., and Putzer, H. (2008) mRNA processing by RNases J1 and J2 affects Bacillus subtilis gene expression on a global scale. Mol Microbiol 70, 183-196. PubMed
- Commichau, F. M., Rothe, F. M., Herzberg, C., Wagner, E., Hellwig, D., Lehnik-Habrink, M., Hammer, E., Völker, U. & Stülke, J. (2009) Novel activities of glycolytic enzymes in Bacillus subtilis: Interactions with essential proteins involved in mRNA processing. Mol. Cell. Proteomics in press PubMed
- Author1, Author2 & Author3 (year) Title Journal volume: page-page. PubMed