Difference between revisions of "CheW"
| Line 121: | Line 121: | ||
=References= | =References= | ||
| − | <pubmed>14651647, </pubmed> | + | <pubmed>,8169224,1601874,14731274,1624415,14651647, </pubmed> |
# Gerth et al. (2008) Clp-dependent proteolysis down-regulates central metabolic pathways in glucose-starved Bacillus subtilis. J Bacteriol 190:321-331 [http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=+17981983 PubMed] | # Gerth et al. (2008) Clp-dependent proteolysis down-regulates central metabolic pathways in glucose-starved Bacillus subtilis. J Bacteriol 190:321-331 [http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=+17981983 PubMed] | ||
# Author1, Author2 & Author3 (year) Title ''Journal'' '''volume:''' page-page. [http://www.ncbi.nlm.nih.gov/sites/entrez/PMID PubMed] | # Author1, Author2 & Author3 (year) Title ''Journal'' '''volume:''' page-page. [http://www.ncbi.nlm.nih.gov/sites/entrez/PMID PubMed] | ||
Revision as of 17:26, 13 June 2009
- Description: modulation of CheA activity in response to attractants, subject to strong Clp-dependent proteolysis upon glucose starvation
| Gene name | cheW |
| Synonyms | |
| Essential | no |
| Product | CheA modulator |
| Function | control of CheA activity |
| MW, pI | 17 kDa, 4.422 |
| Gene length, protein length | 468 bp, 156 aa |
| Immediate neighbours | cheA, cheC |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 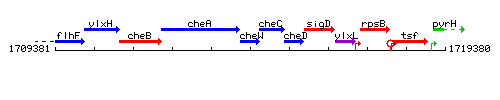
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU16440
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization: cytoplasm (according to Swiss-Prot)
Database entries
- Structure:
- Swiss prot entry: P39802
- KEGG entry: [3]
- E.C. number:
Additional information
- subject to Clp-dependent proteolysis upon glucose starvation PubMed
Expression and regulation
- Operon:
- Additional information: subject to Clp-dependent proteolysis upon glucose starvation PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Michael W Bunn, George W Ordal
Receptor conformational changes enhance methylesterase activity during chemotaxis by Bacillus subtilis.
Mol Microbiol: 2004, 51(3);721-8
[PubMed:14731274]
[WorldCat.org]
[DOI]
(P p)
Virginie Molle, Masaya Fujita, Shane T Jensen, Patrick Eichenberger, José E González-Pastor, Jun S Liu, Richard Losick
The Spo0A regulon of Bacillus subtilis.
Mol Microbiol: 2003, 50(5);1683-701
[PubMed:14651647]
[WorldCat.org]
[DOI]
(P p)
M M Rosario, K L Fredrick, G W Ordal, J D Helmann
Chemotaxis in Bacillus subtilis requires either of two functionally redundant CheW homologs.
J Bacteriol: 1994, 176(9);2736-9
[PubMed:8169224]
[WorldCat.org]
[DOI]
(P p)
D W Hanlon, P B Carpenter, G W Ordal
Influence of attractants and repellents on methyl group turnover on methyl-accepting chemotaxis proteins of Bacillus subtilis and role of CheW.
J Bacteriol: 1992, 174(13);4218-22
[PubMed:1624415]
[WorldCat.org]
[DOI]
(P p)
D W Hanlon, L M Márquez-Magaña, P B Carpenter, M J Chamberlin, G W Ordal
Sequence and characterization of Bacillus subtilis CheW.
J Biol Chem: 1992, 267(17);12055-60
[PubMed:1601874]
[WorldCat.org]
(P p)