Abh
- Description: transcriptional regulator of transition state genes
| Gene name | abh |
| Synonyms | ylxT, yzaA |
| Essential | no |
| Product | transcriptional regulator |
| Function | regulation of gene expression during the transition from growth to stationary phase |
| Gene expression levels in SubtiExpress: abh | |
| Interactions involving this protein in SubtInteract: Abh | |
| MW, pI | 10 kDa, 5.753 |
| Gene length, protein length | 276 bp, 92 aa |
| Immediate neighbours | mreBH, kinC |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 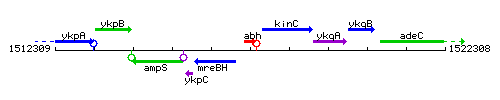
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed
| |
Contents
Categories containing this gene/protein
transition state regulators, cell envelope stress proteins (controlled by SigM, V, W, X, Y)
This gene is a member of the following regulons
The Abh regulon
The gene
Basic information
- Locus tag: BSU14480
Phenotypes of a mutant
- inactivation of abh results in sensitivity against beta-lactam antibiotics that can be restored by induction of slrR expression or by inactivation of the genes encoding major autolysins (lytC, lytF) PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
Genes controlled by Abh
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure: 2FY9 (N-terminal DNA recognition domain)
- UniProt: P39758
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Mark Strauch, Baltimore, USA homepage
Your additional remarks
References
The Abh regulon: PubMed
Other original publications
Yun Luo, John D Helmann
Analysis of the role of Bacillus subtilis σ(M) in β-lactam resistance reveals an essential role for c-di-AMP in peptidoglycan homeostasis.
Mol Microbiol: 2012, 83(3);623-39
[PubMed:22211522]
[WorldCat.org]
[DOI]
(I p)
Veronica Guariglia-Oropeza, John D Helmann
Bacillus subtilis σ(V) confers lysozyme resistance by activation of two cell wall modification pathways, peptidoglycan O-acetylation and D-alanylation of teichoic acids.
J Bacteriol: 2011, 193(22);6223-32
[PubMed:21926231]
[WorldCat.org]
[DOI]
(I p)
Ewan J Murray, Nicola R Stanley-Wall
The sensitivity of Bacillus subtilis to diverse antimicrobial compounds is influenced by Abh.
Arch Microbiol: 2010, 192(12);1059-67
[PubMed:20844865]
[WorldCat.org]
[DOI]
(I p)
Ewan J Murray, Mark A Strauch, Nicola R Stanley-Wall
SigmaX is involved in controlling Bacillus subtilis biofilm architecture through the AbrB homologue Abh.
J Bacteriol: 2009, 191(22);6822-32
[PubMed:19767430]
[WorldCat.org]
[DOI]
(I p)
Yun Luo, John D Helmann
Extracytoplasmic function sigma factors with overlapping promoter specificity regulate sublancin production in Bacillus subtilis.
J Bacteriol: 2009, 191(15);4951-8
[PubMed:19465659]
[WorldCat.org]
[DOI]
(I p)
Mark A Strauch, Benjamin G Bobay, John Cavanagh, Fude Yao, Angelo Wilson, Yoann Le Breton
Abh and AbrB control of Bacillus subtilis antimicrobial gene expression.
J Bacteriol: 2007, 189(21);7720-32
[PubMed:17720793]
[WorldCat.org]
[DOI]
(P p)
Benjamin G Bobay, Geoffrey A Mueller, Richele J Thompson, Alexey G Murzin, Ronald A Venters, Mark A Strauch, John Cavanagh
NMR structure of AbhN and comparison with AbrBN: FIRST insights into the DNA binding promiscuity and specificity of AbrB-like transition state regulator proteins.
J Biol Chem: 2006, 281(30);21399-21409
[PubMed:16702211]
[WorldCat.org]
[DOI]
(P p)
Masakuni Serizawa, Keisuke Kodama, Hiroki Yamamoto, Kazuo Kobayashi, Naotake Ogasawara, Junichi Sekiguchi
Functional analysis of the YvrGHb two-component system of Bacillus subtilis: identification of the regulated genes by DNA microarray and northern blot analyses.
Biosci Biotechnol Biochem: 2005, 69(11);2155-69
[PubMed:16306698]
[WorldCat.org]
[DOI]
(P p)
X Huang, J D Helmann
Identification of target promoters for the Bacillus subtilis sigma X factor using a consensus-directed search.
J Mol Biol: 1998, 279(1);165-73
[PubMed:9636707]
[WorldCat.org]
[DOI]
(P p)
J R LeDeaux, A D Grossman
Isolation and characterization of kinC, a gene that encodes a sensor kinase homologous to the sporulation sensor kinases KinA and KinB in Bacillus subtilis.
J Bacteriol: 1995, 177(1);166-75
[PubMed:8002614]
[WorldCat.org]
[DOI]
(P p)