AlsD
- Description: acetolactate decarboxylase
| Gene name | alsD |
| Synonyms | |
| Essential | no |
| Product | acetolactate decarboxylase) |
| Function | overflow metabolism |
| Metabolic function and regulation of this protein in SubtiPathways: Central C-metabolism | |
| MW, pI | 28 kDa, 4.603 |
| Gene length, protein length | 765 bp, 255 aa |
| Immediate neighbours | ywrO, alsS |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 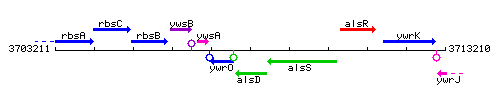
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU36000
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: (2S)-2-hydroxy-2-methyl-3-oxobutanoate = (3R)-3-hydroxybutan-2-one + CO2 (according to Swiss-Prot)
- Protein family: alpha-acetolactate decarboxylase family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification: phosphorylated on ser/ thr/ tyr PubMed
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization:
Database entries
- Structure:
- Swiss prot entry: Q04777
- KEGG entry: [3]
- E.C. number: 4.1.1.5
Additional information
Expression and regulation
Note: since acetate formation requires ackA activation by CcpA there is an indirect effect of CcpA on the alsS-alsD operon: the operon is not expressed in ccpA mutants
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Takashi Inaoka, Takenori Satomura, Yasutaro Fujita, Kozo Ochi
Novel gene regulation mediated by overproduction of secondary metabolite neotrehalosadiamine in Bacillus subtilis.
FEMS Microbiol Lett: 2009, 291(2);151-6
[PubMed:19087206]
[WorldCat.org]
[DOI]
(I p)
Alain Lévine, Françoise Vannier, Cédric Absalon, Lauriane Kuhn, Peter Jackson, Elaine Scrivener, Valérie Labas, Joëlle Vinh, Patrick Courtney, Jérôme Garin, Simone J Séror
Analysis of the dynamic Bacillus subtilis Ser/Thr/Tyr phosphoproteome implicated in a wide variety of cellular processes.
Proteomics: 2006, 6(7);2157-73
[PubMed:16493705]
[WorldCat.org]
[DOI]
(P p)
Heike Reents, Richard Münch, Thorben Dammeyer, Dieter Jahn, Elisabeth Härtig
The Fnr regulon of Bacillus subtilis.
J Bacteriol: 2006, 188(3);1103-12
[PubMed:16428414]
[WorldCat.org]
[DOI]
(P p)
A J Turinsky, T R Moir-Blais, F J Grundy, T M Henkin
Bacillus subtilis ccpA gene mutants specifically defective in activation of acetoin biosynthesis.
J Bacteriol: 2000, 182(19);5611-4
[PubMed:10986270]
[WorldCat.org]
[DOI]
(P p)
M C Renna, N Najimudin, L R Winik, S A Zahler
Regulation of the Bacillus subtilis alsS, alsD, and alsR genes involved in post-exponential-phase production of acetoin.
J Bacteriol: 1993, 175(12);3863-75
[PubMed:7685336]
[WorldCat.org]
[DOI]
(P p)
- Lévine et al. (2006) Analysis of the dynamic Bacillus subtilis Ser/Thr/Tyr phosphoproteome implicated in a wide variety of cellular processes. Proteomics 6: 2157-2173 PubMed
- Renna, M. C., Najimudin, N., Winik, L. R., and Zahler, S. A. (1993). Regulation of the Bacillus subtilis alsS, alsD, and alsR genes involved in post-exponential-phase production of acetoin. J. Bacteriol. 175, 3863-3875. PubMed
- Reents, H., R. Münch, T. Dammeyer, D. Jahn, and E. Härtig. 2006. The Fnr regulon of Bacillus subtilis. J. Bacteriol. 188: 1103-1112. PubMed
- Author1, Author2 & Author3 (year) Title Journal volume: page-page. PubMed