AdhR
- Description: MerR/NmlR-family transcriptional activator of adhA-yraA, responsive to formaldehyde and methylglyoxal
| Gene name | adhR |
| Synonyms | yraB |
| Essential | no |
| Product | MerR/NmlR-family transcriptional activator |
| Function | regulation of the protective response to formaldehyde and methylglyoxal |
| MW, pI | 16 kDa, 9.637 |
| Gene length, protein length | 420 bp, 140 aa |
| Immediate neighbours | yraD, adhA |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 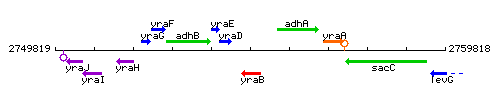
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
transcription factors and their control, resistance against oxidative and electrophile stress
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU27000
Phenotypes of a mutant
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:MerR/NmlR-family of regulators
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification: activity probably redox-controlled via thiol-(S)-alkylation at Cys-52 by aldehydes PubMed
- Cofactor(s):
- Effectors of protein activity:
- Localization: cytoplasmic
Database entries
- Structure:
- UniProt: O06008
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Operon:
- Sigma factor:
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Haike Antelmann,University of Greifswald, Germany
Your additional remarks
References
Reviews
Alastair G McEwan, Karrera Y Djoko, Nathan H Chen, Rafael L M Couñago, Stephen P Kidd, Adam J Potter, Michael P Jennings
Novel bacterial MerR-like regulators their role in the response to carbonyl and nitrosative stress.
Adv Microb Physiol: 2011, 58;1-22
[PubMed:21722790]
[WorldCat.org]
[DOI]
(I p)
Antelmann H, Helmann JD. Thiol-based redox switches and gene regulation. Antioxid Redox Signal. 2011,14:1049-63. PubMed
Original articles
Thi Thu Huyen Nguyen, Warawan Eiamphungporn, Ulrike Mäder, Manuel Liebeke, Michael Lalk, Michael Hecker, John D Helmann, Haike Antelmann
Genome-wide responses to carbonyl electrophiles in Bacillus subtilis: control of the thiol-dependent formaldehyde dehydrogenase AdhA and cysteine proteinase YraA by the MerR-family regulator YraB (AdhR).
Mol Microbiol: 2009, 71(4);876-94
[PubMed:19170879]
[WorldCat.org]
[DOI]
(I p)