Difference between revisions of "CtaG"
| Line 1: | Line 1: | ||
| − | * '''Description:''' formation of functional | + | * '''Description:''' formation of functional cytochrome C-oxidase (caa3) <br/><br/> |
{| align="right" border="1" cellpadding="2" | {| align="right" border="1" cellpadding="2" | ||
| Line 12: | Line 12: | ||
|style="background:#ABCDEF;" align="center"| '''Product''' || unknown | |style="background:#ABCDEF;" align="center"| '''Product''' || unknown | ||
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"|'''Function''' || formation of functional | + | |style="background:#ABCDEF;" align="center"|'''Function''' || formation of functional cytochrome C-oxidase (caa3) |
|- | |- | ||
|style="background:#ABCDEF;" align="center"| '''MW, pI''' || 33 kDa, 6.582 | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || 33 kDa, 6.582 | ||
| Line 88: | Line 88: | ||
=Expression and regulation= | =Expression and regulation= | ||
| − | * '''Operon:''' | + | ** '''Operon:''' ''[[ctaB]]-[[ctaC]]-[[ctaD]]-[[ctaE]]-[[ctaF]]-[[ctaG]]'' {{PubMed|9829923}} |
* '''[[Sigma factor]]:''' | * '''[[Sigma factor]]:''' | ||
* '''Regulation:''' | * '''Regulation:''' | ||
| + | ** expressed under anaerobic conditions ([[ResD]]) {{PubMed|9829923}} | ||
* '''Regulatory mechanism:''' | * '''Regulatory mechanism:''' | ||
| + | ** [[ResD]]: transcription activation {{PubMed|9829923}} | ||
* '''Additional information:''' | * '''Additional information:''' | ||
| Line 118: | Line 120: | ||
=References= | =References= | ||
| − | <pubmed>14766920,, </pubmed> | + | <pubmed>14766920,9829923, </pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 20:23, 8 October 2009
- Description: formation of functional cytochrome C-oxidase (caa3)
| Gene name | ctaG |
| Synonyms | |
| Essential | no |
| Product | unknown |
| Function | formation of functional cytochrome C-oxidase (caa3) |
| MW, pI | 33 kDa, 6.582 |
| Gene length, protein length | 891 bp, 297 aa |
| Immediate neighbours | ctaF, ylbA |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 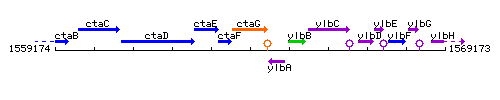
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU14930
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization: cell membrane (according to Swiss-Prot)
Database entries
- Structure:
- UniProt: O34329
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Jenny Bengtsson, Claes von Wachenfeldt, Lena Winstedt, Per Nygaard, Lars Hederstedt
CtaG is required for formation of active cytochrome c oxidase in Bacillus subtilis.
Microbiology (Reading): 2004, 150(Pt 2);415-425
[PubMed:14766920]
[WorldCat.org]
[DOI]
(P p)
X Liu, H W Taber
Catabolite regulation of the Bacillus subtilis ctaBCDEF gene cluster.
J Bacteriol: 1998, 180(23);6154-63
[PubMed:9829923]
[WorldCat.org]
[DOI]
(P p)