DeaD
- Description: DEAD-box RNA helicase
| Gene name | deaD |
| Synonyms | yxiN |
| Essential | no |
| Product | ATP-dependent RNA helicase |
| Function | RNA helicase |
| MW, pI | 53 kDa, 7.706 |
| Gene length, protein length | 1437 bp, 479 aa |
| Immediate neighbours | yxiO, yxiM |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 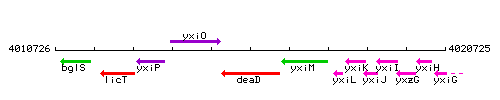
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU39110
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: RNA helicase
- Protein family: DEAD-box RNA helicase
Extended information on the protein
- Kinetic information:
- Domains:
- Helicase domain 1: AS 1-203
- Linker region: AS 204-212
- Helicase domain 2: AS 207-368
- RNA-binding domain: AS 404-479
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization:
Database entries
- UniProt: P42305
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant: GP1121 (tet), available in the Stülke lab
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Stülke lab
- Antibody:
Labs working on this gene/protein
Dagmar Klostermeier, Biozentrum Basel, Switzerland homepage
Your additional remarks
References
Regula Aregger, Dagmar Klostermeier
The DEAD box helicase YxiN maintains a closed conformation during ATP hydrolysis.
Biochemistry: 2009, 48(45);10679-81
[PubMed:19839642]
[WorldCat.org]
[DOI]
(I p)
Anne R Karow, Dagmar Klostermeier
A conformational change in the helicase core is necessary but not sufficient for RNA unwinding by the DEAD box helicase YxiN.
Nucleic Acids Res: 2009, 37(13);4464-71
[PubMed:19474341]
[WorldCat.org]
[DOI]
(I p)
Bettina Theissen, Anne R Karow, Jürgen Köhler, Airat Gubaev, Dagmar Klostermeier
Cooperative binding of ATP and RNA induces a closed conformation in a DEAD box RNA helicase.
Proc Natl Acad Sci U S A: 2008, 105(2);548-53
[PubMed:18184816]
[WorldCat.org]
[DOI]
(I p)
Anne R Karow, Bettina Theissen, Dagmar Klostermeier
Authentic interdomain communication in an RNA helicase reconstituted by expressed protein ligation of two helicase domains.
FEBS J: 2007, 274(2);463-73
[PubMed:17229151]
[WorldCat.org]
[DOI]
(P p)
Jonathan M Caruthers, YaoXiong Hu, David B McKay
Structure of the second domain of the Bacillus subtilis DEAD-box RNA helicase YxiN.
Acta Crystallogr Sect F Struct Biol Cryst Commun: 2006, 62(Pt 12);1191-5
[PubMed:17142894]
[WorldCat.org]
[DOI]
(I p)
Shuying Wang, Yaoxiong Hu, Michael T Overgaard, Fedor V Karginov, Olke C Uhlenbeck, David B McKay
The domain of the Bacillus subtilis DEAD-box helicase YxiN that is responsible for specific binding of 23S rRNA has an RNA recognition motif fold.
RNA: 2006, 12(6);959-67
[PubMed:16611943]
[WorldCat.org]
[DOI]
(P p)
Fedor V Karginov, Jonathan M Caruthers, YaoXiong Hu, David B McKay, Olke C Uhlenbeck
YxiN is a modular protein combining a DEx(D/H) core and a specific RNA-binding domain.
J Biol Chem: 2005, 280(42);35499-505
[PubMed:16118224]
[WorldCat.org]
[DOI]
(P p)