Difference between revisions of "DesR"
| (6 intermediate revisions by 2 users not shown) | |||
| Line 14: | Line 14: | ||
|style="background:#ABCDEF;" align="center"|'''Function''' || regulation of cold shock expression of des | |style="background:#ABCDEF;" align="center"|'''Function''' || regulation of cold shock expression of des | ||
|- | |- | ||
| − | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http:// | + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU19200 desR] |
|- | |- | ||
| − | |colspan="2" style="background:#FAF8CC;" align="center"| '''Interactions involving this protein in [http:// | + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Interactions involving this protein in [http://subtiwiki.uni-goettingen.de/interact/ ''Subt''Interact]''': [http://subtiwiki.uni-goettingen.de/interact/index.php?protein=DesR DesR] |
|- | |- | ||
| − | |colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: <br/>[http://subtiwiki.uni-goettingen.de/ | + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: <br/>[http://subtiwiki.uni-goettingen.de/subtipathways/search.php?enzyme=desR desR]''' |
|- | |- | ||
|style="background:#ABCDEF;" align="center"| '''MW, pI''' || 22 kDa, 4.885 | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || 22 kDa, 4.885 | ||
| Line 26: | Line 26: | ||
|style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[desK]]'', ''[[yocH]]'' | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[desK]]'', ''[[yocH]]'' | ||
|- | |- | ||
| − | | | + | |style="background:#FAF8CC;" align="center"|'''Sequences'''||[http://bsubcyc.org/BSUB/sequence-aa?type=GENE&object=BSU19200 Protein] [http://bsubcyc.org/BSUB/sequence?type=GENE&object=BSU19200 DNA] [http://bsubcyc.org/BSUB/seq-selector?chromosome=CHROM-1&object=BSU19200 DNA_with_flanks] |
|- | |- | ||
|colspan="2" | '''Genetic context''' <br/> [[Image:yocG_context.gif]] | |colspan="2" | '''Genetic context''' <br/> [[Image:yocG_context.gif]] | ||
<div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | <div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | ||
|- | |- | ||
| − | |colspan="2" |'''[http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=desR_2091705_2092304_1 Expression at a glance]'''   {{PubMed|22383849}}<br/>[[Image:desR_expression.png|500px]] | + | |colspan="2" |'''[http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=desR_2091705_2092304_1 Expression at a glance]'''   {{PubMed|22383849}}<br/>[[Image:desR_expression.png|500px|link=http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU19200]] |
|- | |- | ||
|} | |} | ||
| Line 39: | Line 39: | ||
<br/><br/><br/><br/> | <br/><br/><br/><br/> | ||
<br/><br/><br/><br/> | <br/><br/><br/><br/> | ||
| − | + | <br/><br/> | |
| − | |||
| − | |||
| − | |||
| − | |||
= [[Categories]] containing this gene/protein = | = [[Categories]] containing this gene/protein = | ||
| Line 65: | Line 61: | ||
=== Database entries === | === Database entries === | ||
| + | * '''BsubCyc:''' [http://bsubcyc.org/BSUB/NEW-IMAGE?type=NIL&object=BSU19200&redirect=T BSU19200] | ||
* '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/yocFG.html] | * '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/yocFG.html] | ||
| Line 71: | Line 68: | ||
=== Additional information=== | === Additional information=== | ||
| − | |||
| − | |||
| − | |||
=The protein= | =The protein= | ||
| Line 103: | Line 97: | ||
=== Database entries === | === Database entries === | ||
| + | * '''BsubCyc:''' [http://bsubcyc.org/BSUB/NEW-IMAGE?type=NIL&object=BSU19200&redirect=T BSU19200] | ||
* '''Structure:''' | * '''Structure:''' | ||
| Line 150: | Line 145: | ||
=References= | =References= | ||
| − | <pubmed>10094672,12399512, 19595746, 11285232, 17087771 12207704</pubmed> | + | <pubmed>10094672,12399512, 19595746, 11285232, 17087771 12207704 25406381 </pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Latest revision as of 07:02, 24 November 2014
- Description: two-component response regulator, regulation of cold shock expression of des
| Gene name | desR |
| Synonyms | yocG |
| Essential | no |
| Product | two-component response regulator |
| Function | regulation of cold shock expression of des |
| Gene expression levels in SubtiExpress: desR | |
| Interactions involving this protein in SubtInteract: DesR | |
| Metabolic function and regulation of this protein in SubtiPathways: desR | |
| MW, pI | 22 kDa, 4.885 |
| Gene length, protein length | 597 bp, 199 aa |
| Immediate neighbours | desK, yocH |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 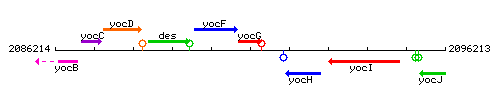
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed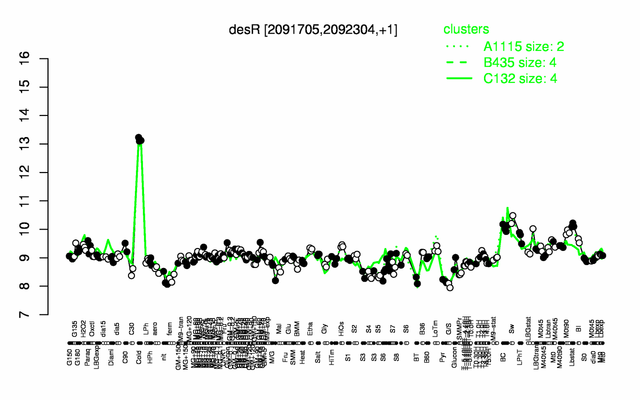
| |
Contents
Categories containing this gene/protein
lipid metabolism/ other, transcription factors and their control, cold stress proteins, membrane proteins, phosphoproteins
This gene is a member of the following regulons
The DesR regulon:
The gene
Basic information
- Locus tag: BSU19200
Phenotypes of a mutant
Database entries
- BsubCyc: BSU19200
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: transcription activation of the des operon when phosphorylated by DesK
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification: phosphorylated by DesK on an Asp residue
- Cofactor(s):
- Effectors of protein activity:
- Localization: cell membrane (according to Swiss-Prot)
Database entries
- BsubCyc: BSU19200
- Structure:
- UniProt: O34723
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Sigma factor:
- Regulation:
- induced by cold shock (18-fold) PubMed
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
- Diego de Mendoza, Universidad Nacional de Rosario, Argentine homepage
Your additional remarks
References
Felipe Trajtenberg, Daniela Albanesi, Natalia Ruétalo, Horacio Botti, Ariel E Mechaly, Marcos Nieves, Pablo S Aguilar, Larisa Cybulski, Nicole Larrieux, Diego de Mendoza, Alejandro Buschiazzo
Allosteric activation of bacterial response regulators: the role of the cognate histidine kinase beyond phosphorylation.
mBio: 2014, 5(6);e02105
[PubMed:25406381]
[WorldCat.org]
[DOI]
(I e)
Sebastián R Najle, María E Inda, Diego de Mendoza, Larisa E Cybulski
Oligomerization of Bacillus subtilis DesR is required for fine tuning regulation of membrane fluidity.
Biochim Biophys Acta: 2009, 1790(10);1238-43
[PubMed:19595746]
[WorldCat.org]
[DOI]
(P p)
Pablo S Aguilar, Diego de Mendoza
Control of fatty acid desaturation: a mechanism conserved from bacteria to humans.
Mol Microbiol: 2006, 62(6);1507-14
[PubMed:17087771]
[WorldCat.org]
[DOI]
(P p)
Carsten L Beckering, Leif Steil, Michael H W Weber, Uwe Völker, Mohamed A Marahiel
Genomewide transcriptional analysis of the cold shock response in Bacillus subtilis.
J Bacteriol: 2002, 184(22);6395-402
[PubMed:12399512]
[WorldCat.org]
[DOI]
(P p)
Larisa E Cybulski, Daniela Albanesi, María C Mansilla, Silvia Altabe, Pablo S Aguilar, Diego de Mendoza
Mechanism of membrane fluidity optimization: isothermal control of the Bacillus subtilis acyl-lipid desaturase.
Mol Microbiol: 2002, 45(5);1379-88
[PubMed:12207704]
[WorldCat.org]
[DOI]
(P p)
P S Aguilar, A M Hernandez-Arriaga, L E Cybulski, A C Erazo, D de Mendoza
Molecular basis of thermosensing: a two-component signal transduction thermometer in Bacillus subtilis.
EMBO J: 2001, 20(7);1681-91
[PubMed:11285232]
[WorldCat.org]
[DOI]
(P p)
C Fabret, V A Feher, J A Hoch
Two-component signal transduction in Bacillus subtilis: how one organism sees its world.
J Bacteriol: 1999, 181(7);1975-83
[PubMed:10094672]
[WorldCat.org]
[DOI]
(P p)