Difference between revisions of "FusA"
| Line 1: | Line 1: | ||
| − | + | ||
{| align="right" border="1" cellpadding="2" | {| align="right" border="1" cellpadding="2" | ||
| Line 13: | Line 13: | ||
|- | |- | ||
|style="background:#ABCDEF;" align="center"|'''Function''' || translation | |style="background:#ABCDEF;" align="center"|'''Function''' || translation | ||
| + | |- | ||
| + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://cellpublisher.gobics.de/subtiexpress/ ''Subti''Express]''': [http://cellpublisher.gobics.de/subtiexpress/bsu/BSU01120 fusA] | ||
|- | |- | ||
|style="background:#ABCDEF;" align="center"| '''MW, pI''' || 76 kDa, 4.615 | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || 76 kDa, 4.615 | ||
Revision as of 14:22, 6 August 2012
| Gene name | fusA |
| Synonyms | fus |
| Essential | yes PubMed |
| Product | elongation factor G |
| Function | translation |
| Gene expression levels in SubtiExpress: fusA | |
| MW, pI | 76 kDa, 4.615 |
| Gene length, protein length | 2076 bp, 692 aa |
| Immediate neighbours | rpsG, tufA |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 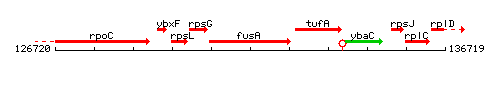
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed
| |
Contents
Categories containing this gene/protein
translation, essential genes, phosphoproteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU01120
Phenotypes of a mutant
essential PubMed
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: hydrolyses GTP
- Protein family: EF-G/EF-2 subfamily (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification: phosphorylation on Ser-213 AND Ser-302 AND Ser-569 AND Ser-680 AND (Thr-24 OR Thr-25) AND (Thr-43 OR Ser 48) PubMed, PubMed
- Cofactor(s):
- Effectors of protein activity:
- Localization: cytoplasm (according to Swiss-Prot)
Database entries
- UniProt: P80868
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Operon:
- Sigma factor:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Nina Clementi, Anna Chirkova, Barbara Puffer, Ronald Micura, Norbert Polacek
Atomic mutagenesis reveals A2660 of 23S ribosomal RNA as key to EF-G GTPase activation.
Nat Chem Biol: 2010, 6(5);344-51
[PubMed:20348921]
[WorldCat.org]
[DOI]
(I p)
Christine Eymann, Dörte Becher, Jörg Bernhardt, Katrin Gronau, Anja Klutzny, Michael Hecker
Dynamics of protein phosphorylation on Ser/Thr/Tyr in Bacillus subtilis.
Proteomics: 2007, 7(19);3509-26
[PubMed:17726680]
[WorldCat.org]
[DOI]
(P p)
Boris Macek, Ivan Mijakovic, Jesper V Olsen, Florian Gnad, Chanchal Kumar, Peter R Jensen, Matthias Mann
The serine/threonine/tyrosine phosphoproteome of the model bacterium Bacillus subtilis.
Mol Cell Proteomics: 2007, 6(4);697-707
[PubMed:17218307]
[WorldCat.org]
[DOI]
(P p)
A AEvarsson, E Brazhnikov, M Garber, J Zheltonosova, Y Chirgadze, S al-Karadaghi, L A Svensson, A Liljas
Three-dimensional structure of the ribosomal translocase: elongation factor G from Thermus thermophilus.
EMBO J: 1994, 13(16);3669-77
[PubMed:8070397]
[WorldCat.org]
[DOI]
(P p)