Difference between revisions of "GltC"
(→Biological materials) |
(→Expression and regulation) |
||
| Line 91: | Line 91: | ||
=Expression and regulation= | =Expression and regulation= | ||
| − | * '''Operon:''' ''[[gltC]]'' | + | * '''Operon:''' ''[[gltC]]'' (according to [http://dbtbs.hgc.jp/COG/prom/gltC.html DBTBS]) |
| − | * '''Sigma factor:''' [[SigA]] | + | * '''Sigma factor:''' [[SigA]] {{PubMed|2548995}} |
| − | * '''Regulation:''' autoregulation by GltC | + | * '''Regulation:''' autoregulation by GltC {{PubMed|2548995}} |
| − | * '''Regulatory mechanism:''' autorepression | + | * '''Regulatory mechanism:''' autorepression {{PubMed|2548995}} |
* '''Database entries:''' [http://dbtbs.hgc.jp/COG/prom/gltC.html DBTBS] | * '''Database entries:''' [http://dbtbs.hgc.jp/COG/prom/gltC.html DBTBS] | ||
Revision as of 20:31, 18 October 2009
- Description: Transcriptional activator of the gltAB operon. Activates expression of the operon in the absence of arginine.
| Gene name | gltC |
| Synonyms | |
| Essential | No |
| Product | transcriptional regulator (LysR family) |
| Function | positive regulation of the glutamate synthase operon (gltAB) |
| Metabolic function and regulation of this protein in SubtiPathways: Ammonium/ glutamate | |
| MW, pI | 33.9 kDa, 5.62 |
| Gene length, protein length | 900 bp, 300 amino acids |
| Immediate neighbours | gltA, proJ |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 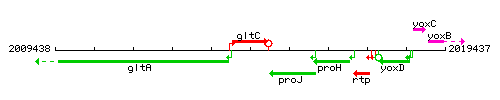
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU18460
Phenotypes of a mutant
gltC mutants are auxotrophic for glutamate.
Database entries
- DBTBS entry: [1]
- SubtiList entry:[2]
Additional information
The protein
Basic information/ Evolution
- Protein family: glutamate synthase family (according to Swiss-Prot) LysR-type transcription regulator PubMed
- Paralogous protein(s): none, but there are 19 members of the LysR family in B. subtilis
Extended information on the protein
- Kinetic information:
- Domains: DNA-binding helix-turn-helix motif: AA 18 ... 37
- Modification:
- Cofactor(s):
- Effectors of protein activity: 2-oxoglutarate stimulates transcription activation, glutamate inhibits transcription activation PubMed
- Interactions:
- Localization:
Database entries
- Structure:
- UniProt: P20668
- KEGG entry: [3]
Additional information
Expression and regulation
- Regulation: autoregulation by GltC PubMed
- Regulatory mechanism: autorepression PubMed
- Database entries: DBTBS
- Additional information:
Biological materials
- Mutant: GP344 (erm), GP738 (spc) (available in Stülke lab)
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Stülke lab
- Antibody: available in Stülke lab
Labs working on this gene/protein
Linc Sonenshein, Tufts University, Boston, MA, USA Homepage
Jörg Stülke, University of Göttingen, Germany Homepage
Your additional remarks
References