Difference between revisions of "QueF"
| (22 intermediate revisions by 6 users not shown) | |||
| Line 1: | Line 1: | ||
| − | * '''Description:''' synthesis of the modified | + | * '''Description:''' nitrile reductase (NADPH-dependent 7-cyano-7-deazaguanine reductase), synthesis of the modified ribonucleotide queuosine <br/><br/> |
{| align="right" border="1" cellpadding="2" | {| align="right" border="1" cellpadding="2" | ||
| Line 10: | Line 10: | ||
|style="background:#ABCDEF;" align="center"| '''Essential''' || no | |style="background:#ABCDEF;" align="center"| '''Essential''' || no | ||
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Product''' || | + | |style="background:#ABCDEF;" align="center"| '''Product''' || nitrile reductase |
|- | |- | ||
|style="background:#ABCDEF;" align="center"|'''Function''' || tRNA modification | |style="background:#ABCDEF;" align="center"|'''Function''' || tRNA modification | ||
| + | |- | ||
| + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU13750 queF] | ||
|- | |- | ||
|style="background:#ABCDEF;" align="center"| '''MW, pI''' || 19 kDa, 4.927 | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || 19 kDa, 4.927 | ||
| Line 20: | Line 22: | ||
|style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[queE]]'', ''[[ykvN]]'' | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[queE]]'', ''[[ykvN]]'' | ||
|- | |- | ||
| − | | | + | |style="background:#FAF8CC;" align="center"|'''Sequences'''||[http://bsubcyc.org/BSUB/sequence-aa?type=GENE&object=BSU13750 Protein] [http://bsubcyc.org/BSUB/sequence?type=GENE&object=BSU13750 DNA] [http://bsubcyc.org/BSUB/seq-selector?chromosome=CHROM-1&object=BSU13750 DNA_with_flanks] |
|- | |- | ||
|colspan="2" | '''Genetic context''' <br/> [[Image:ykvM_context.gif]] | |colspan="2" | '''Genetic context''' <br/> [[Image:ykvM_context.gif]] | ||
<div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | <div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | ||
| + | |- | ||
| + | |colspan="2" |'''[http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=queF_1441291_1441788_1 Expression at a glance]'''   {{PubMed|22383849}}<br/>[[Image:queF_expression.png|500px|link=http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU13750]] | ||
|- | |- | ||
|} | |} | ||
__TOC__ | __TOC__ | ||
| + | <br/><br/><br/><br/> | ||
| + | <br/><br/><br/><br/> | ||
| + | <br/><br/><br/><br/> | ||
| + | <br/><br/><br/><br/> | ||
| + | <br/><br/><br/><br/> | ||
| + | |||
<br/><br/> | <br/><br/> | ||
| + | |||
| + | = [[Categories]] containing this gene/protein = | ||
| + | {{SubtiWiki category|[[translation]]}} | ||
| + | |||
| + | = This gene is a member of the following [[regulons]] = | ||
| + | {{SubtiWiki regulon|[[preQ1 riboswitch]]}} | ||
=The gene= | =The gene= | ||
| Line 40: | Line 56: | ||
=== Database entries === | === Database entries === | ||
| + | * '''BsubCyc:''' [http://bsubcyc.org/BSUB/NEW-IMAGE?type=NIL&object=BSU13750&redirect=T BSU13750] | ||
* '''DBTBS entry:''' no entry | * '''DBTBS entry:''' no entry | ||
| Line 46: | Line 63: | ||
=== Additional information=== | === Additional information=== | ||
| + | |||
| + | |||
| Line 64: | Line 83: | ||
* '''Domains:''' | * '''Domains:''' | ||
| − | * '''Modification:''' | + | * '''Modification:''' active site Cys56 is S-bacillithiolated by NaOCl stress {{PubMed|22938038}} |
* '''Cofactor(s):''' | * '''Cofactor(s):''' | ||
| Line 70: | Line 89: | ||
* '''Effectors of protein activity:''' | * '''Effectors of protein activity:''' | ||
| − | * '''Interactions:''' | + | * '''[[SubtInteract|Interactions]]:''' |
| − | * '''Localization:''' cytoplasm (according to Swiss-Prot) | + | * '''[[Localization]]:''' cytoplasm (according to Swiss-Prot) |
=== Database entries === | === Database entries === | ||
| + | * '''BsubCyc:''' [http://bsubcyc.org/BSUB/NEW-IMAGE?type=NIL&object=BSU13750&redirect=T BSU13750] | ||
| − | * '''Structure:''' | + | * '''Structure:''' [http://www.rcsb.org/pdb/explore/explore.do?structureId=4F8b 4F8B] (covalent thioimide intermediate of the unimodular nitrile reductase [[QueF]]) {{PubMed|22787148}} |
| − | * ''' | + | * '''UniProt:''' [http://www.uniprot.org/uniprot/O31678 O31678] |
| − | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu | + | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu:BSU13750] |
* '''E.C. number:''' [http://www.expasy.org/enzyme/1.7.1.13 1.7.1.13] | * '''E.C. number:''' [http://www.expasy.org/enzyme/1.7.1.13 1.7.1.13] | ||
| Line 88: | Line 108: | ||
=Expression and regulation= | =Expression and regulation= | ||
| − | * '''Operon:''' | + | * '''Operon:''' ''[[queC]]-[[queD]]-[[queE]]-[[queF]]'' {{PubMed|14660578}} |
| + | |||
| + | * '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=queF_1441291_1441788_1 queF] {{PubMed|22383849}} | ||
| − | * ''' | + | * '''Sigma factor:''' |
* '''Regulation:''' | * '''Regulation:''' | ||
| + | ** repressed in the presence of queuosine ([[preQ1 riboswitch]]) {{PubMed|19285444}} | ||
* '''Regulatory mechanism:''' | * '''Regulatory mechanism:''' | ||
| + | ** [[preQ1 riboswitch]]: transcriptional antitermination in the absence of queuosine {{PubMed|19285444}} | ||
| − | * '''Additional information:''' | + | * '''Additional information:''' |
| + | ** number of protein molecules per cell (minimal medium with glucose and ammonium): 303 {{PubMed|24696501}} | ||
| + | ** number of protein molecules per cell (complex medium with amino acids, without glucose): 913 {{PubMed|24696501}} | ||
=Biological materials = | =Biological materials = | ||
| Line 118: | Line 144: | ||
=References= | =References= | ||
| − | <pubmed>14660578,, </pubmed> | + | <pubmed>22938038,14660578,15767583 16511203 ,17929836 19285444 17384645 22787148 </pubmed> |
| − | + | ||
| + | [[Category:Protein-coding genes]] | ||
Latest revision as of 09:43, 17 April 2014
- Description: nitrile reductase (NADPH-dependent 7-cyano-7-deazaguanine reductase), synthesis of the modified ribonucleotide queuosine
| Gene name | queF |
| Synonyms | ykvM |
| Essential | no |
| Product | nitrile reductase |
| Function | tRNA modification |
| Gene expression levels in SubtiExpress: queF | |
| MW, pI | 19 kDa, 4.927 |
| Gene length, protein length | 495 bp, 165 aa |
| Immediate neighbours | queE, ykvN |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 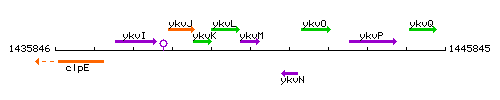
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed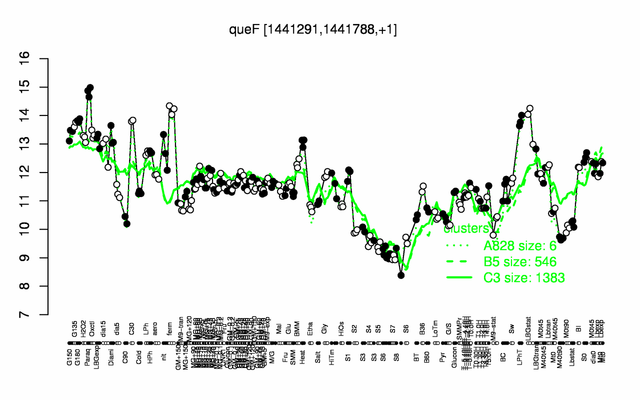
| |
Contents
Categories containing this gene/protein
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU13750
Phenotypes of a mutant
Database entries
- BsubCyc: BSU13750
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: 7-aminomethyl-7-carbaguanine + 2 NADP+ = 7-cyano-7-carbaguanine + 2 NADPH (according to Swiss-Prot)
- Protein family: QueF type 1 subfamily (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification: active site Cys56 is S-bacillithiolated by NaOCl stress PubMed
- Cofactor(s):
- Effectors of protein activity:
- Localization: cytoplasm (according to Swiss-Prot)
Database entries
- BsubCyc: BSU13750
- UniProt: O31678
- KEGG entry: [2]
- E.C. number: 1.7.1.13
Additional information
Expression and regulation
- Sigma factor:
- Regulation:
- repressed in the presence of queuosine (preQ1 riboswitch) PubMed
- Regulatory mechanism:
- preQ1 riboswitch: transcriptional antitermination in the absence of queuosine PubMed
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Bui Khanh Chi, Alexandra A Roberts, Tran Thi Thanh Huyen, Katrin Bäsell, Dörte Becher, Dirk Albrecht, Chris J Hamilton, Haike Antelmann
S-bacillithiolation protects conserved and essential proteins against hypochlorite stress in firmicutes bacteria.
Antioxid Redox Signal: 2013, 18(11);1273-95
[PubMed:22938038]
[WorldCat.org]
[DOI]
(I p)
Vimbai M Chikwana, Boguslaw Stec, Bobby W K Lee, Valérie de Crécy-Lagard, Dirk Iwata-Reuyl, Manal A Swairjo
Structural basis of biological nitrile reduction.
J Biol Chem: 2012, 287(36);30560-70
[PubMed:22787148]
[WorldCat.org]
[DOI]
(I p)
Mijeong Kang, Robert Peterson, Juli Feigon
Structural Insights into riboswitch control of the biosynthesis of queuosine, a modified nucleotide found in the anticodon of tRNA.
Mol Cell: 2009, 33(6);784-90
[PubMed:19285444]
[WorldCat.org]
[DOI]
(I p)
Bobby W K Lee, Steven G Van Lanen, Dirk Iwata-Reuyl
Mechanistic studies of Bacillus subtilis QueF, the nitrile oxidoreductase involved in queuosine biosynthesis.
Biochemistry: 2007, 46(44);12844-54
[PubMed:17929836]
[WorldCat.org]
[DOI]
(P p)
Adam Roth, Wade C Winkler, Elizabeth E Regulski, Bobby W K Lee, Jinsoo Lim, Inbal Jona, Jeffrey E Barrick, Ankita Ritwik, Jane N Kim, Rüdiger Welz, Dirk Iwata-Reuyl, Ronald R Breaker
A riboswitch selective for the queuosine precursor preQ1 contains an unusually small aptamer domain.
Nat Struct Mol Biol: 2007, 14(4);308-17
[PubMed:17384645]
[WorldCat.org]
[DOI]
(P p)
Manal A Swairjo, Robert R Reddy, Bobby Lee, Steven G Van Lanen, Shannon Brown, Valérie de Crécy-Lagard, Dirk Iwata-Reuyl, Paul Schimmel
Crystallization and preliminary X-ray characterization of the nitrile reductase QueF: a queuosine-biosynthesis enzyme.
Acta Crystallogr Sect F Struct Biol Cryst Commun: 2005, 61(Pt 10);945-8
[PubMed:16511203]
[WorldCat.org]
[DOI]
(I p)
Steven G Van Lanen, John S Reader, Manal A Swairjo, Valérie de Crécy-Lagard, Bobby Lee, Dirk Iwata-Reuyl
From cyclohydrolase to oxidoreductase: discovery of nitrile reductase activity in a common fold.
Proc Natl Acad Sci U S A: 2005, 102(12);4264-9
[PubMed:15767583]
[WorldCat.org]
[DOI]
(P p)
John S Reader, David Metzgar, Paul Schimmel, Valérie de Crécy-Lagard
Identification of four genes necessary for biosynthesis of the modified nucleoside queuosine.
J Biol Chem: 2004, 279(8);6280-5
[PubMed:14660578]
[WorldCat.org]
[DOI]
(P p)