Difference between revisions of "Sandbox"
| Line 1: | Line 1: | ||
| − | * '''Description:''' DNA-Gyrase (subunit | + | * '''Description:''' DNA-Gyrase (subunit A) <br/><br/> |
{| align="right" border="1" cellpadding="2" | {| align="right" border="1" cellpadding="2" | ||
|- | |- | ||
|style="background:#ABCDEF;" align="center"|'''Gene name''' | |style="background:#ABCDEF;" align="center"|'''Gene name''' | ||
| − | |'' | + | |''gyrA'' |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Synonyms''' || '' | + | |style="background:#ABCDEF;" align="center"| '''Synonyms''' || ''nalA '' |
|- | |- | ||
|style="background:#ABCDEF;" align="center"| '''Essential''' || yes [http://www.ncbi.nlm.nih.gov/pubmed/12682299 PubMed] | |style="background:#ABCDEF;" align="center"| '''Essential''' || yes [http://www.ncbi.nlm.nih.gov/pubmed/12682299 PubMed] | ||
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Product''' || DNA-Gyrase (subunit | + | |style="background:#ABCDEF;" align="center"| '''Product''' || DNA-Gyrase (subunit A) |
|- | |- | ||
|style="background:#ABCDEF;" align="center"|'''Function''' || DNA supercoiling, <br/>initation of replication cycle and DNA elongation | |style="background:#ABCDEF;" align="center"|'''Function''' || DNA supercoiling, <br/>initation of replication cycle and DNA elongation | ||
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || | + | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || 91 kDa, 5.215 |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Gene length, protein length''' || | + | |style="background:#ABCDEF;" align="center"| '''Gene length, protein length''' || 2463 bp, 821 aa |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[ | + | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[gyrB]]'', ''[[rrnO-16S]]'' |
|- | |- | ||
| − | |colspan="2" style="background:#FAF8CC;" align="center"|'''Get the DNA and protein [http://srs.ebi.ac.uk/srsbin/cgi-bin/wgetz?-e+[EMBLCDS: | + | |colspan="2" style="background:#FAF8CC;" align="center"|'''Get the DNA and protein [http://srs.ebi.ac.uk/srsbin/cgi-bin/wgetz?-e+[EMBLCDS:CAB11783]+-newId sequences] <br/> (Barbe ''et al.'', 2009)''' |
|- | |- | ||
| − | |colspan="2" | '''Genetic context''' <br/> [[Image: | + | |colspan="2" | '''Genetic context''' <br/> [[Image:GyrA_context.png]] |
<div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | <div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | ||
|- | |- | ||
| Line 35: | Line 35: | ||
=== Basic information === | === Basic information === | ||
| − | * '''Locus tag:''' | + | * '''Locus tag:''' BSU00070 |
===Phenotypes of a mutant === | ===Phenotypes of a mutant === | ||
| Line 43: | Line 43: | ||
=== Database entries === | === Database entries === | ||
| − | * '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/ | + | * '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/gyrA.html] |
| − | * '''SubtiList entry:''' [http://genolist.pasteur.fr/SubtiList/genome.cgi?gene_detail+ | + | * '''SubtiList entry:''' [http://genolist.pasteur.fr/SubtiList/genome.cgi?gene_detail+BG10071] |
=== Additional information=== | === Additional information=== | ||
| Line 56: | Line 56: | ||
* '''Catalyzed reaction/ biological activity:''' ATP-dependent breakage, passage and rejoining of double-stranded DNA (according to Swiss-Prot) | * '''Catalyzed reaction/ biological activity:''' ATP-dependent breakage, passage and rejoining of double-stranded DNA (according to Swiss-Prot) | ||
| − | * '''Protein family:''' | + | * '''Protein family:''' topoisomerase gyrA/parC subunit family (according to Swiss-Prot) |
| − | * '''Paralogous protein(s):''' [[ | + | * '''Paralogous protein(s):''' [[ParC]] |
=== Extended information on the protein === | === Extended information on the protein === | ||
| Line 74: | Line 74: | ||
* '''Interactions:''' [[GyrA]]-[[GyrB]] | * '''Interactions:''' [[GyrA]]-[[GyrB]] | ||
| − | * '''Localization:''' nucleoid-associated in actively growing cells [http://www.ncbi.nlm.nih.gov/sites/entrez/9539793 PubMed] | + | * '''Localization:''' Nucleoid (Homogeneous) [http://www.ncbi.nlm.nih.gov/sites/entrez/16479537 PubMed] nucleoid-associated in actively growing cells [http://www.ncbi.nlm.nih.gov/sites/entrez/9539793 PubMed] |
=== Database entries === | === Database entries === | ||
| Line 80: | Line 80: | ||
* '''Structure:''' | * '''Structure:''' | ||
| − | * '''Swiss prot entry:''' [http://www.uniprot.org/uniprot/ | + | * '''Swiss prot entry:''' [http://www.uniprot.org/uniprot/P05653 P05653] |
| − | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu+ | + | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu+BSU00070] |
* '''E.C. number:''' | * '''E.C. number:''' | ||
=== Additional information=== | === Additional information=== | ||
| + | |||
| + | :* subject to Clp-dependent proteolysis upon glucose starvation [http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=+17981983 PubMed] | ||
=Expression and regulation= | =Expression and regulation= | ||
| − | * '''Operon:''' '' | + | * '''Operon:''' ''gyrA'' [http://www.ncbi.nlm.nih.gov/sites/entrez/2987848 PubMed] |
* '''[[Sigma factor]]:''' [[SigA]] [http://www.ncbi.nlm.nih.gov/sites/entrez/2987848 PubMed] | * '''[[Sigma factor]]:''' [[SigA]] [http://www.ncbi.nlm.nih.gov/sites/entrez/2987848 PubMed] | ||
| Line 98: | Line 100: | ||
* '''Regulatory mechanism:''' | * '''Regulatory mechanism:''' | ||
| − | * '''Additional information:''' | + | * '''Additional information:''' subject to Clp-dependent proteolysis upon glucose starvation [http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=+17981983 PubMed], GyrA is subject to Clp-dependent proteolysis upon glucose starvation [http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=+17981983 PubMed] |
=Biological materials = | =Biological materials = | ||
| Line 122: | Line 124: | ||
=References= | =References= | ||
| − | <pubmed>17981983 ,9539793 , 2987848 </pubmed> | + | <pubmed>16479537,17981983 ,9539793 , 2987848 </pubmed> |
Revision as of 12:47, 14 June 2009
- Description: DNA-Gyrase (subunit A)
| Gene name | gyrA |
| Synonyms | nalA |
| Essential | yes PubMed |
| Product | DNA-Gyrase (subunit A) |
| Function | DNA supercoiling, initation of replication cycle and DNA elongation |
| MW, pI | 91 kDa, 5.215 |
| Gene length, protein length | 2463 bp, 821 aa |
| Immediate neighbours | gyrB, rrnO-16S |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 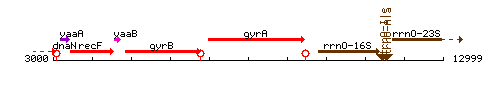
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU00070
Phenotypes of a mutant
essential PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: ATP-dependent breakage, passage and rejoining of double-stranded DNA (according to Swiss-Prot)
- Protein family: topoisomerase gyrA/parC subunit family (according to Swiss-Prot)
- Paralogous protein(s): ParC
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure:
- Swiss prot entry: P05653
- KEGG entry: [3]
- E.C. number:
Additional information
- subject to Clp-dependent proteolysis upon glucose starvation PubMed
Expression and regulation
- Operon: gyrA PubMed
- Regulation:
- Regulatory mechanism:
- Additional information: subject to Clp-dependent proteolysis upon glucose starvation PubMed, GyrA is subject to Clp-dependent proteolysis upon glucose starvation PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Dagmar Klostermeier, Biozentrum Basel, Switzerland homepage
Your additional remarks
References
Ulf Gerth, Holger Kock, Ilja Kusters, Stephan Michalik, Robert L Switzer, Michael Hecker
Clp-dependent proteolysis down-regulates central metabolic pathways in glucose-starved Bacillus subtilis.
J Bacteriol: 2008, 190(1);321-31
[PubMed:17981983]
[WorldCat.org]
[DOI]
(I p)
Jean-Christophe Meile, Ling Juan Wu, S Dusko Ehrlich, Jeff Errington, Philippe Noirot
Systematic localisation of proteins fused to the green fluorescent protein in Bacillus subtilis: identification of new proteins at the DNA replication factory.
Proteomics: 2006, 6(7);2135-46
[PubMed:16479537]
[WorldCat.org]
[DOI]
(P p)
W M Huang, J L Libbey, P van der Hoeven, S X Yu
Bipolar localization of Bacillus subtilis topoisomerase IV, an enzyme required for chromosome segregation.
Proc Natl Acad Sci U S A: 1998, 95(8);4652-7
[PubMed:9539793]
[WorldCat.org]
[DOI]
(P p)
N Ogasawara, S Moriya, H Yoshikawa
Structure and function of the region of the replication origin of the Bacillus subtilis chromosome. IV. Transcription of the oriC region and expression of DNA gyrase genes and other open reading frames.
Nucleic Acids Res: 1985, 13(7);2267-79
[PubMed:2987848]
[WorldCat.org]
[DOI]
(P p)