Difference between revisions of "Spo0A"
(→Other original Publications) |
|||
| Line 14: | Line 14: | ||
|style="background:#ABCDEF;" align="center"|'''Function''' || initiation of sporulation | |style="background:#ABCDEF;" align="center"|'''Function''' || initiation of sporulation | ||
|- | |- | ||
| − | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http:// | + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU24220 spo0A] |
|- | |- | ||
|colspan="2" style="background:#FAF8CC;" align="center"| '''Interactions involving this protein in [http://cellpublisher.gobics.de/subtinteract/startpage/start/ ''Subt''Interact]''': [http://cellpublisher.gobics.de/subtinteract/interactionList/2/Spo0A Spo0A] | |colspan="2" style="background:#FAF8CC;" align="center"| '''Interactions involving this protein in [http://cellpublisher.gobics.de/subtinteract/startpage/start/ ''Subt''Interact]''': [http://cellpublisher.gobics.de/subtinteract/interactionList/2/Spo0A Spo0A] | ||
| Line 26: | Line 26: | ||
|style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[yqiG]]'', ''[[spoIVB]]'' | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[yqiG]]'', ''[[spoIVB]]'' | ||
|- | |- | ||
| − | | | + | |style="background:#FAF8CC;" align="center"|'''Sequences'''||[http://bsubcyc.org/BSUB/sequence-aa?type=GENE&object=BSU24220 Protein] [http://bsubcyc.org/BSUB/sequence?type=GENE&object=BSU24220 DNA] [http://bsubcyc.org/BSUB/seq-selector?chromosome=CHROM-1&object=BSU24220 Advanced_DNA] |
|- | |- | ||
|colspan="2" | '''Genetic context''' <br/> [[Image:spo0A_context.gif]] | |colspan="2" | '''Genetic context''' <br/> [[Image:spo0A_context.gif]] | ||
Revision as of 13:15, 13 May 2013
- Description: phosphorelay regulator, initiation of sporulation, coordinates DNA replication and initiation of sporulation by binding to sites close to the oriC
| Gene name | spo0A |
| Synonyms | spo0C, spo0G, spoIIL, sof-1 |
| Essential | no |
| Product | phosphorelay response regulator |
| Function | initiation of sporulation |
| Gene expression levels in SubtiExpress: spo0A | |
| Interactions involving this protein in SubtInteract: Spo0A | |
| Metabolic function and regulation of this protein in SubtiPathways: Biofilm, Nucleotides (regulation), Ammonium/ glutamate, Central C-metabolism,Sugar catabolism, Phosphorelay, Stress, tRNA charging,Lipid synthesis, Protein secretion | |
| MW, pI | 29 kDa, 5.989 |
| Gene length, protein length | 801 bp, 267 aa |
| Immediate neighbours | yqiG, spoIVB |
| Sequences | Protein DNA Advanced_DNA |
Genetic context 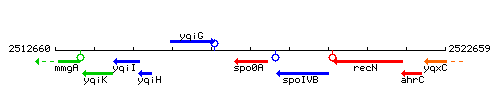
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed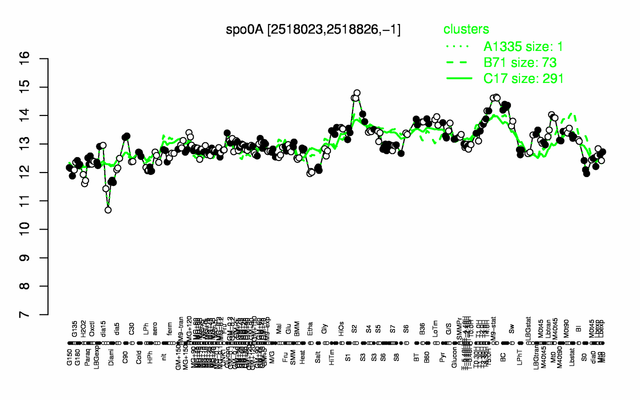
| |
Contents
Categories containing this gene/protein
transcription factors and their control, phosphorelay, biofilm formation, phosphoproteins
This gene is a member of the following regulons
The Spo0A regulon
The gene
Basic information
- Locus tag: BSU24220
Phenotypes of a mutant
- inactivation of spo0A restores beta-lactam resistance in a sigM mutant PubMed
- altered cell death pattern in colonies PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- activation of gene expression at the onset of sporulation (Spo0A regulon)
- coordination between DNA replication and initiation of sporulation by binding to sites close to the oriC (this limits re-initiation of replication) PubMed
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification: receives phosphorylation from Spo0B, dephosphorylation by Spo0E, direct phosphorylation by KinC upon potassium leakage PubMed
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure: 1QMP (with phosphorylated aspartate, Geobacillus stearothermophilus), 1FC3 (trans-activation domain, Geobacillus stearothermophilus), 1LQ1 (complex with DNA)
- UniProt: P06534
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Operon: spo0A PubMed
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
- Tony Wilkinson, York University, U.K. homepage
- Imrich Barak, Slovak Academy of Science, Bratislava, Slovakia homepage
- Charles Moran, Emory University, NC, USA homepage
Your additional remarks
References
Reviews
The Spo0A regulon
Masaya Fujita, José Eduardo González-Pastor, Richard Losick
High- and low-threshold genes in the Spo0A regulon of Bacillus subtilis.
J Bacteriol: 2005, 187(4);1357-68
[PubMed:15687200]
[WorldCat.org]
[DOI]
(P p)
Virginie Molle, Masaya Fujita, Shane T Jensen, Patrick Eichenberger, José E González-Pastor, Jun S Liu, Richard Losick
The Spo0A regulon of Bacillus subtilis.
Mol Microbiol: 2003, 50(5);1683-701
[PubMed:14651647]
[WorldCat.org]
[DOI]
(P p)
Other original Publications
Additional publications: PubMed